dna structure and replication hl
Nature of science:
Making careful observations—Rosalind Franklin’s X-ray diffraction provided crucial evidence that DNA is a double helix. (1.8)
Understandings:
• Nucleosomes help to supercoil the DNA.
• DNA structure suggested a mechanism for DNA replication.
• DNA polymerases can only add nucleotides to the 3’ end of a primer.
• DNA replication is continuous on the leading strand and discontinuous on the lagging strand.
• DNA replication is carried out by a complex system of enzymes.
• Some regions of DNA do not code for proteins but have other important functions.
Making careful observations—Rosalind Franklin’s X-ray diffraction provided crucial evidence that DNA is a double helix. (1.8)
Understandings:
• Nucleosomes help to supercoil the DNA.
• DNA structure suggested a mechanism for DNA replication.
• DNA polymerases can only add nucleotides to the 3’ end of a primer.
• DNA replication is continuous on the leading strand and discontinuous on the lagging strand.
• DNA replication is carried out by a complex system of enzymes.
• Some regions of DNA do not code for proteins but have other important functions.
DNA structure suggested a mechanism for DNA replication
Several lines of experimental evidence came together to lead to the knowledge of the structure of DNA: molecular modelling pionered by Linus pauling, X ray diffraction patterns discerned from the careful photogrpahs of Rosalind Franklin.
One of the Watson and crick's first model had the sugar phosphate strands wrapped around one another with the nitrogen bases facing outwards. Roselyn countered this model given that the bases were hydrophobic n comparison of the sugar phosphate. After many other models, the key factors was discovered, the complementary pairing rules, once the new model was proposed the complementary base pairing suggested a mechanism by which DNA replication could occur - one of the key requirements that any structural model would have to address. The watson-crick model led to the hypothesis of semi conservation replication.
One of the Watson and crick's first model had the sugar phosphate strands wrapped around one another with the nitrogen bases facing outwards. Roselyn countered this model given that the bases were hydrophobic n comparison of the sugar phosphate. After many other models, the key factors was discovered, the complementary pairing rules, once the new model was proposed the complementary base pairing suggested a mechanism by which DNA replication could occur - one of the key requirements that any structural model would have to address. The watson-crick model led to the hypothesis of semi conservation replication.
The role of nucleosomes in dna packing
NUCLEOSOMES HELP TO SUPERCOIL DNA
One difference between eukaryotics DNA and bacterial DNA is that eukaryotic DNA is associated with proteins called histones. Most groups of prokaryotes have DNA that is not associated with histones, or proteins like histones. for this reason, prokaryotic DNA is referred to being naked
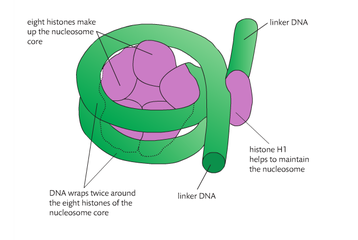
Histones are used by the cell to package the DNA into structures called nucleosomes. A nucleosome consists of a central core of eight histones proteins with DNA coiled around the proteins. The 8 proteins or octamer, consist of 2 copies of 4 different types of histones. A short section of linker DNA connects one nucleosome to the next. An additional histone protein molecule called H1, serves to bind the DNA to the core particle.
The association of histones with DNA contributes to a pattern know as supercoiling. An analogy is of you twist an elastic band repeatedly eventually it forms an additional pattern of coils. Supercoiling allows a great length of DNA to be packed into much smaller space within the nucleus. The nucleosome is an adaptation that facilitates the packing of the large genomes that the eukaryotes possess. The H1 histones bids in such a way to form a structure called the 30nm fiber tat facilitates further packing
One difference between eukaryotics DNA and bacterial DNA is that eukaryotic DNA is associated with proteins called histones. Most groups of prokaryotes have DNA that is not associated with histones, or proteins like histones. for this reason, prokaryotic DNA is referred to being naked
Histones are used by the cell to package the DNA into structures called nucleosomes. A nucleosome consists of a central core of eight histones proteins with DNA coiled around the proteins. The 8 proteins or octamer, consist of 2 copies of 4 different types of histones. A short section of linker DNA connects one nucleosome to the next. An additional histone protein molecule called H1, serves to bind the DNA to the core particle.
The association of histones with DNA contributes to a pattern know as supercoiling. An analogy is of you twist an elastic band repeatedly eventually it forms an additional pattern of coils. Supercoiling allows a great length of DNA to be packed into much smaller space within the nucleus. The nucleosome is an adaptation that facilitates the packing of the large genomes that the eukaryotes possess. The H1 histones bids in such a way to form a structure called the 30nm fiber tat facilitates further packing
the leading strand and the lagging strand
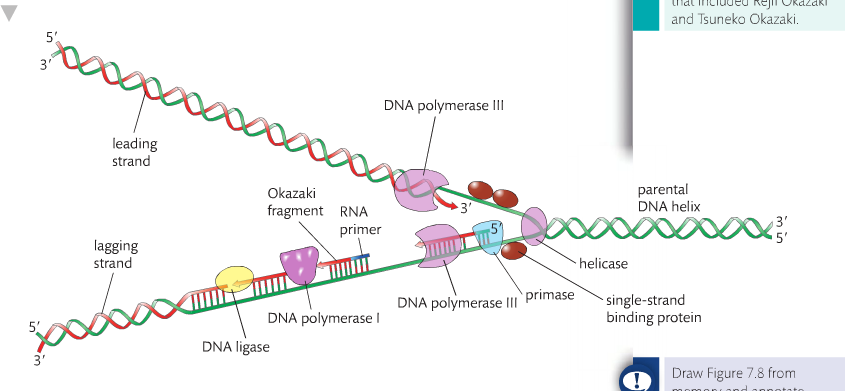
Because the 2 strands of the DNA double helix are arranged in an antiparallel fashion, synthesis on the 2 strands occurs in very different ways. One strand, the leading strand, is made continuously following the fork as it opens. The other strand known as the lagging strand, is made in fragments moving away from the replication fork. New fragments are created on the lagging strand as the replication fork exposes more of the template strand. These fragments are called the Okazaki fragments
proteins involved in replication
DNA replication is carried out by a complex system of enzymes.
Replication involves the formation and movement of the replication fork and synthesis of the leading and lagging strands. Proteins are involved as enzymes at each stage but also serve a number of other functions.
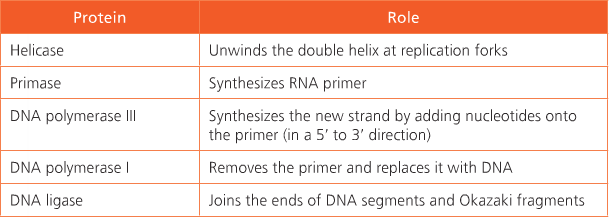
The enzyme helicase unwinds the DNA at the replication fork and the enzyme topoisomerase the strain that develops ahead o the helicase. Single-stranded binding proteins keep the strands apart long enough to allow the template strand to be copied. Starting replication requires an RNA primer. Note that on the lagging strand there are a number of primer but there is jut one on the leading strand. The enzyme DN primase creates one RNA primer on the leading strand and many RNA primers on the lagging strand. The RNA primer is necessary to initiate the activity of DNA polymerase.
DNA polymerase is responsible for covalently linking the deoxyribonucleotide monophosphate to the 3’ end of the growing strand. Different functions such as proof-reading polymeraziation and removal of RNA primers once they are no longer needed.
Replication involves the formation and movement of the replication fork and synthesis of the leading and lagging strands. Proteins are involved as enzymes at each stage but also serve a number of other functions.
The enzyme helicase unwinds the DNA at the replication fork and the enzyme topoisomerase the strain that develops ahead o the helicase. Single-stranded binding proteins keep the strands apart long enough to allow the template strand to be copied. Starting replication requires an RNA primer. Note that on the lagging strand there are a number of primer but there is jut one on the leading strand. The enzyme DN primase creates one RNA primer on the leading strand and many RNA primers on the lagging strand. The RNA primer is necessary to initiate the activity of DNA polymerase.
DNA polymerase is responsible for covalently linking the deoxyribonucleotide monophosphate to the 3’ end of the growing strand. Different functions such as proof-reading polymeraziation and removal of RNA primers once they are no longer needed.
the direction of replication
DNa polymerase can only add nucleotides to the 3’ end if a primer.
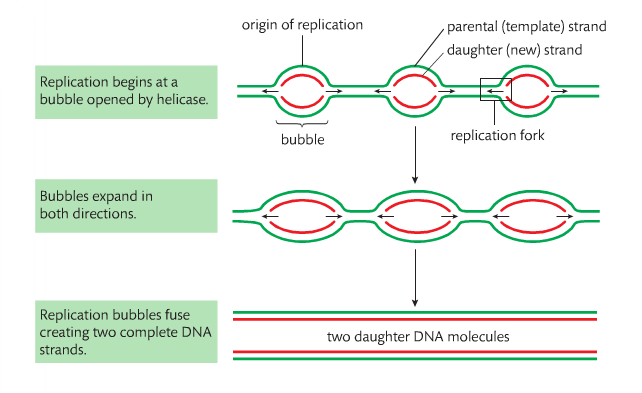
Within DNA molecules, DNA replication begins at sites called origins of replication. In prokaryotes there is one site and in eukaryotes there are many. Replication occurs in both directions away from the origin. The result appears as a replication bubble in electron micrographs.
The phosphate egorup of new DNA nucleoties is added to the 3’carbon of the deoxyribose of the nucleotide as the end of the chain. Replication therefore occurs in the 5’ to 3’ direction.
Within DNA molecules, DNA replication begins at sites called origins of replication. In prokaryotes there is one site and in eukaryotes there are many. Replication occurs in both directions away from the origin. The result appears as a replication bubble in electron micrographs.
The phosphate egorup of new DNA nucleoties is added to the 3’carbon of the deoxyribose of the nucleotide as the end of the chain. Replication therefore occurs in the 5’ to 3’ direction.
non-coding regions of Dna have important functions
Some regions of DNa do not code for proteins but have other important functions.
The cellular machinery operates according to a genetic code. DNA is used as a guide for the production of polypeptides using the genetic code. However, only some DNA sequences code for the production of polypeptides. These are called coding sequences. There are a number of non-coding sequences found in genomes. Some of them have functions such as those sequences that are used as a guide to produce tRNA and rRNA. Some non-coding regions play an important role in the regulation of gene expression such as enhancers and silencers.
Most of the eukaryotic genome is non-coding
Within the genome, especially in eukaryotes, repetitive sequences can be common, Tere are two types of repetitive sequences: moderately repetitive sequences and highly repetitive sequences. Together they form between 5 and 60 per cent of the genome. In humans nearly 60 percent of the DNA is in repetitive sequences.
One such area of repetitive sequences occurs on the end of the eukaryotes chromosomes called telomeres. The telomeres serves a protective function. During interphase, the enzyme that replicates DNA cannot continue replicating all the way to the end of the chromosome. If cells went through the cell cycle without telomeres, they would lose the genes at the end of the chromosomes. Sacrificing the repetitive sequences found in telomeres.
The cellular machinery operates according to a genetic code. DNA is used as a guide for the production of polypeptides using the genetic code. However, only some DNA sequences code for the production of polypeptides. These are called coding sequences. There are a number of non-coding sequences found in genomes. Some of them have functions such as those sequences that are used as a guide to produce tRNA and rRNA. Some non-coding regions play an important role in the regulation of gene expression such as enhancers and silencers.
Most of the eukaryotic genome is non-coding
Within the genome, especially in eukaryotes, repetitive sequences can be common, Tere are two types of repetitive sequences: moderately repetitive sequences and highly repetitive sequences. Together they form between 5 and 60 per cent of the genome. In humans nearly 60 percent of the DNA is in repetitive sequences.
One such area of repetitive sequences occurs on the end of the eukaryotes chromosomes called telomeres. The telomeres serves a protective function. During interphase, the enzyme that replicates DNA cannot continue replicating all the way to the end of the chromosome. If cells went through the cell cycle without telomeres, they would lose the genes at the end of the chromosomes. Sacrificing the repetitive sequences found in telomeres.
Applications and skills:
• Application: Rosalind Franklin’s and Maurice Wilkins’ investigation of DNA structure by X-ray diffraction.
• Application: Use of nucleotides containing dideoxyribonucleic acid to stop DNA replication in preparation of samples for base sequencing.
• Application: Tandem repeats are used in DNA profiling.
• Skill: Analysis of results of the Hershey and Chase experiment providing evidence that DNA is the genetic material.
• Skill: Utilization of molecular visualization software to analyse the association between protein and DNA within a nucleosome.
Guidance:
• Details of DNA replication differ between prokaryotes and eukaryotes. Only the prokaryotic system is expected.
• The proteins and enzymes involved in DNA replication should include helicase, DNA gyrase, single strand binding proteins, DNA primase and DNA polymerases I and III.
• The regions of DNA that do not code for proteins should be limited to regulators of gene expression, introns, telomeres and genes for tRNAs.
Theory of knowledge:
• Highly repetitive sequences were once classified as “junk DNA” showing a degree of confidence that it had no role. To what extent do the labels and categories used in the pursuit of knowledge affect the knowledge we obtain?
The labels that we attribute to things can cause severe damage in the perspective and opinion of that thing that was labelled. It is always better to not label someone or something before you know it is actually true, you can cause severe damage to the feelings of someone if the label is wrong but everyone else doesn't know that. labels can shape or knowledge into the same way that the person that labelled the thing or the person is, which therefore makes you have a biased knowledge. Therefore, we should never label someone or something to the extend that we have to be 100% that the label is truth and that the knowledge we are getting from it is not biased or based in someone else's opinion.